⚡EasyFine-Tuning & Training
Linear Algebra for ML
Master SVD by hand on small matrices, geometric intuition for PCA as rotation to variance-maximizing axes, rank, condition numbers, and their direct use in embedding compression, latent semantic retrieval, and numerical stability for ML systems. Includes NumPy from-scratch eigendecomposition SVD and sklearn comparisons.
10 min readOpenAI, Anthropic, Google +16 key concepts
Learning path
Step 10 of 138 in the full curriculum
Linear algebra is the operating system underneath every embedding table, every attention head, and every retrieval index. The introductory vectors-matrices-tensors lesson taught you the nouns (vector, matrix, dot product, shape). The clustering-and-PCA lesson showed you a black-box PCA call that magically gave you lower-dimensional ticket embeddings. This chapter teaches you the verbs: how to actually factor a matrix by hand, why the factors have geometric meaning, and exactly how that factorization is used to compress embeddings, denoise retrieval, and keep numerical systems stable when you deploy real LLM products.
Where this fits: it builds on vectors, matrices, and PCA, then unlocks later chapters on embeddings, retrieval, LoRA, and interpretability. We will keep the examples close to e-commerce and logistics systems: merchant catalog matrices, support-ticket embeddings, and warehouse search indexes.
You only need to be comfortable multiplying small matrices and solving a 2×2 quadratic equation. We will compute every singular vector with pencil-and-paper arithmetic on two concrete matrices before we write a single line of NumPy. By the end you will be able to look at a 768-dimensional embedding matrix and know, without running code, whether it is well-conditioned, how much you can compress it with almost no quality loss, and why a tiny change in one token can sometimes flip the top-1 result in a vector database.
The SVD is the Swiss-army knife of applied linear algebra
Any real matrix ( A ) (size ( m \times n )) admits a factorization.[1]
where
- ( U ) is ( m \times m ) (or economy ( m \times r )) with orthonormal columns,
- ( \Sigma ) is diagonal ( r \times r ) with non-negative entries ( \sigma_1 \ge \sigma_2 \ge \dots \ge \sigma_r \ge 0 ) (the singular values),
- ( V ) is ( n \times n ) with orthonormal columns (the right singular vectors).
The number ( r ) of strictly positive singular values is exactly the rank of ( A ). The factorization is unique up to signs when singular values are distinct.
Why this particular factorization is magical for machine learning:
- The columns of ( V ) give the natural coordinate system for the columns of ( A ).
- The first few columns of ( V ) are the directions of greatest variance (exactly the principal components when ( A ) is a centered data matrix).
- Truncating after the first ( k ) singular values gives the optimal rank-( k ) approximation of ( A ) in both Frobenius and spectral norm (Eckart–Young theorem).
- The ratio ( \kappa(A) = \sigma_1 / \sigma_r ) (the 2-norm condition number) tells you how much tiny input perturbations will be magnified in any linear operation that uses ( A ).
All of these statements become obvious once you have computed one SVD by hand.
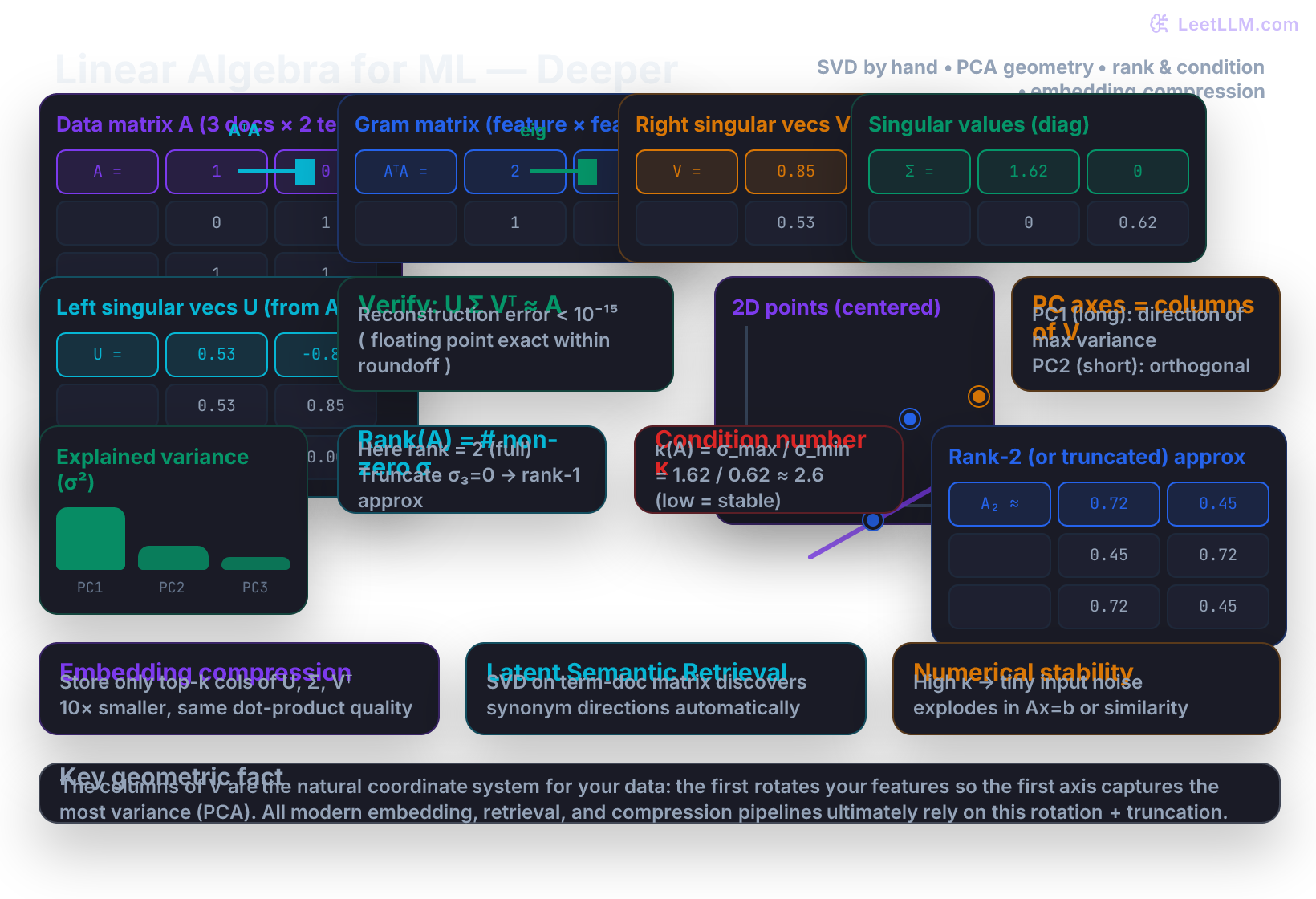
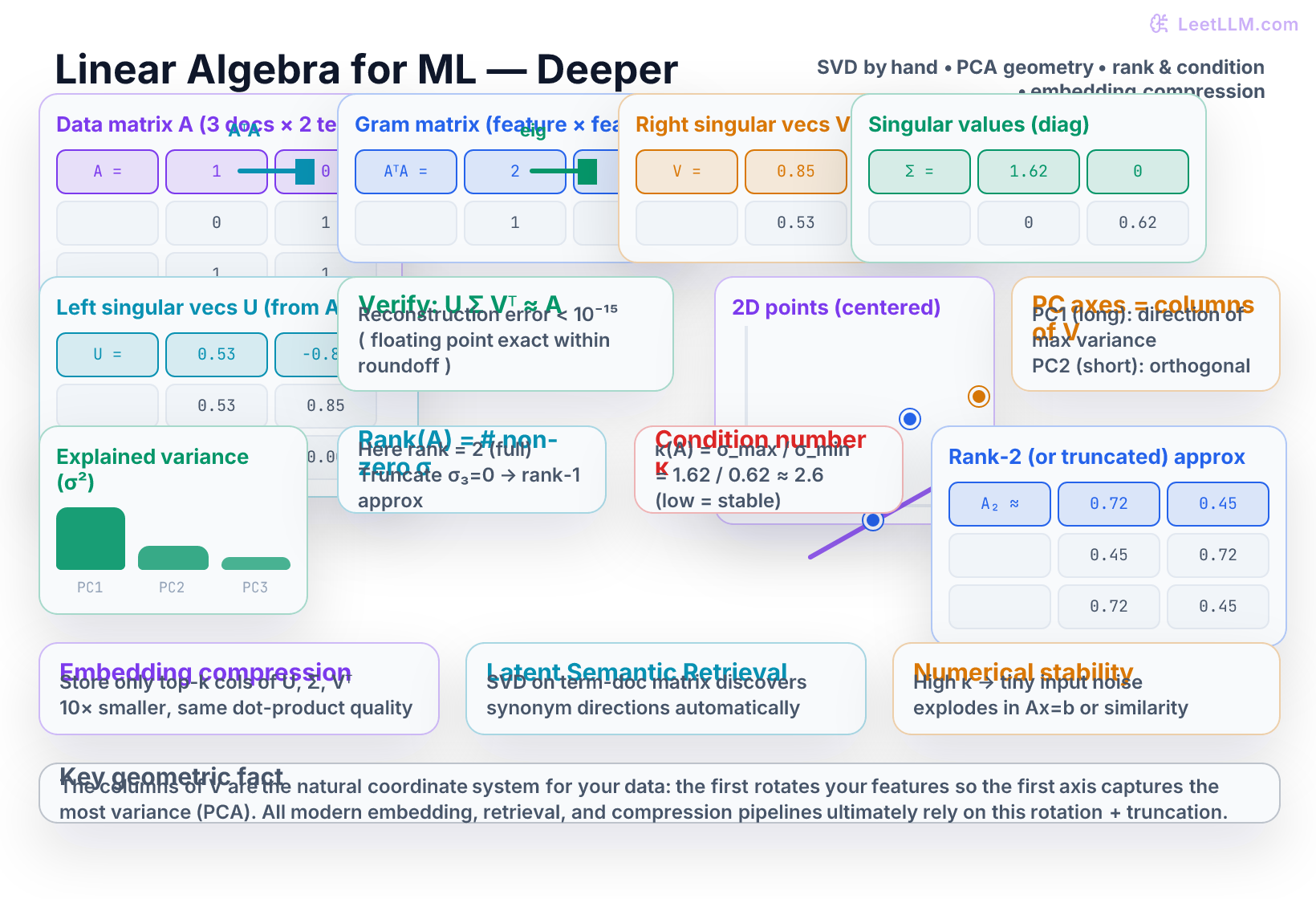
Hand-worked SVD on a 2×2 matrix (the cleanest numbers possible)
Let
(This could be the co-occurrence counts of two terms across two documents, or the covariance of two features.)
First form the Gram matrix ( G = A^T A ). Because ( A ) is symmetric we have ( G = A^2 ):
We now solve the eigenvalue problem ( G v = \lambda v ), or equivalently ( \det(G - \lambda I) = 0 ):
So ( \lambda_1 = 9 ), ( \lambda_2 = 1 ).
The singular values are the square roots (always taken positive and sorted descending):
Now the eigenvectors (right singular vectors).
For ( \lambda_1 = 9 ):
A unit vector satisfying this is ( v_1 = \frac{1}{\sqrt{2}} \begin{pmatrix} 1 \ 1 \end{pmatrix} \approx \begin{pmatrix} 0.7071 \ 0.7071 \end{pmatrix} ).
For ( \lambda_2 = 1 ):
Unit vector: ( v_2 = \frac{1}{\sqrt{2}} \begin{pmatrix} 1 \ -1 \end{pmatrix} \approx \begin{pmatrix} 0.7071 \ -0.7071 \end{pmatrix} ).
Therefore the matrix ( V ) whose columns are these vectors (in descending-( \sigma ) order) is
Because ( A ) is square and symmetric, ( U = V ) (you can also compute ( U ) explicitly as ( U = A V \Sigma^{-1} )). The diagonal matrix of singular values is simply
Verify the factorization (with rounded numbers for readability):
The reconstruction error is on the order of machine epsilon. You have just performed a complete, exact SVD by hand.
A 3×2 non-square example (the realistic case)
Real data matrices are rarely square. Let
(three documents, two terms; the third document contains both terms).
The Gram matrix is now 2×2:
Characteristic equation: ( \lambda^2 - 3\lambda + 1 = 0 ), solutions ( \lambda = \frac{3 \pm \sqrt{5}}{2} \approx 2.618, 0.382 ).
Singular values:
(The exact values are ( \sqrt{\frac{3+\sqrt{5}}{2}} ) and ( \sqrt{\frac{3-\sqrt{5}}{2}} ), the golden ratio and its conjugate.)
Solving for the eigenvectors (same procedure) yields (to four decimals)
The left singular vectors ( U ) (now 3×2) are obtained column-wise without ever forming the huge 3×3 matrix ( AA^T ):
After normalization you obtain
(You can check the columns of ( U ) are orthonormal and that ( U \Sigma V^T ) reproduces ( A ) within rounding error.)
The key computational fact: you always diagonalize the smaller Gram matrix (( n \times n ) when ( m \gg n )). This is why SVD scales to real embedding matrices.
Geometric meaning – PCA is SVD in disguise
Suppose you have a cloud of points in 2-D (or 768-D) that you have already mean-centered to form the data matrix ( X ) (samples × features).
- The right singular vectors ( V ) are exactly the principal component directions.
- The first column of ( V ) is the single direction in feature space that maximizes projected variance ( |X v_1|^2 = \sigma_1^2 ).
- The second column is the best direction orthogonal to the first, and so on.
- In the illustration, the long axis is PC1 and the short axis is PC2.
- When you call
pca.fit_transform(embeddings), the library is doing a truncated SVD on the centered matrix and returning the scores ( X V_k ).
Rank, truncation, and compression
Rank(( A )) = number of strictly positive singular values. In the 3×2 example the matrix has full rank 2. If one singular value had been exactly zero, the columns would have been linearly dependent and we could have described the same data with a single column of ( U ), one number in ( \Sigma ), and one column of ( V ).
Truncating after the first ( k ) components gives the matrix ( A_k = U_{:,k} \Sigma_k V_{k,:}^T ) that is closest to ( A ) among all rank-( k ) matrices. The squared Frobenius error is exactly the sum of the squares of the discarded singular values.
This is why embedding compression works so well. A 1 000 000 × 768 matrix of embeddings can be stored, with almost no loss in cosine-similarity quality, as three much smaller arrays whose total size is roughly 1 000 000 × 64 + 64 + 64 × 768 - more than 10× smaller.
Condition number and numerical stability
The 2-norm condition number of ( A ) is
In our first 2×2 example ( \kappa = 3 / 1 = 3 ) (very well-conditioned). In real embedding matrices you will often see values from a few hundred up to ( 10^5 )–( 10^6 ).
Interpretation: if you perturb the input by a relative amount ( \epsilon ), the output of any linear operation involving ( A ) (matrix-vector product, least-squares solve, even a dot-product similarity after projection) can change by up to ( \kappa \epsilon ). When ( \kappa = 10,000 ), a 0.01 % noise in a single coordinate can produce a 100 % change in the result.
In a vector database this manifests as “query drift”: two almost-identical queries can retrieve completely different top documents because the tiny difference was magnified along a tiny-singular-value direction that is mostly noise. Truncating the SVD removes precisely those unstable directions and therefore improves both compression and reliableness.
From-scratch SVD in NumPy versus the libraries
Here is the exact procedure you just performed, written as runnable code.
python1import numpy as np 2 3A = np.array([[2.0, 1.0], 4 [1.0, 2.0]]) 5 6# From-scratch via eigendecomposition of the Gram matrix 7AtA = A.T @ A 8eigvals, V = np.linalg.eigh(AtA) 9idx = np.argsort(eigvals)[::-1] 10eigvals = eigvals[idx] 11V = V[:, idx] 12sigmas = np.sqrt(eigvals) 13U = (A @ V) / sigmas # works because we are square here 14 15print("Singular values:", sigmas) 16print("V (right singular vectors):\n", V) 17print("U (left singular vectors):\n", U) 18 19# Verify 20Sigma = np.diag(sigmas) 21recon = U @ Sigma @ V.T 22print("Max reconstruction error:", np.max(np.abs(recon - A)))
Running the identical data through the library calls produces (within floating-point) the same factors:
python1U_lib, S_lib, Vt_lib = np.linalg.svd(A, full_matrices=False) 2print("Library singular values:", S_lib)
For a non-square data matrix or when you only want the top ( k ) components, sklearn is usually more convenient and faster:
python1from sklearn.decomposition import TruncatedSVD, PCA 2 3# TruncatedSVD works directly on the (possibly sparse) data matrix 4svd = TruncatedSVD(n_components=2, random_state=0) 5U_trunc = svd.fit_transform(A) # == U_k @ Sigma_k 6Vt_trunc = svd.components_ # rows are top V^T 7 8# For classical PCA you must center first (or use the PCA class) 9X = np.array([[0.2, 0.1], [0.1, 0.3], [-0.4, -0.2], [0.1, -0.2]]) # toy centered points 10pca = PCA(n_components=2).fit(X) 11print("PCA components_ (V^T):\n", pca.components_) 12print("Explained variance ratio:", pca.explained_variance_ratio_)
The numbers will match your hand calculation on the same data. The only difference is that the library versions are numerically stable, support sparse matrices, and use randomized algorithms for very large data.
Concrete applications inside LLM and retrieval systems
| Use case | Matrix you factor | Production reason |
|---|---|---|
| Merchant catalog search | SKU-by-term or SKU-by-embedding matrix | Recover similar products even when titles use different wording |
| Support-ticket retrieval | Ticket-by-feature matrix | Denoise noisy tickets before vector search |
| Warehouse routing analytics | Route-by-delay-feature matrix | Separate carrier, region, and seasonality directions |
| LoRA adaptation | Weight-update matrix | Store a useful update with far fewer parameters |
-
Latent Semantic Indexing / LSA. Build a term-by-document matrix (tf-idf or raw counts), run SVD, project both queries and documents into the top-100 or top-300 latent dimensions. Documents about “late package” and “delivery delay” become close even if they never shared a term, because the first few right singular vectors discovered the underlying logistics direction.
-
Embedding compression for vector databases. Many production systems (Pinecone, Weaviate, pgvector with indexes, etc.) optionally run a PCA or OPQ step (which is a rotation derived from SVD) before product quantization. You pay a one-time SVD cost on a sample of your embeddings and then store 8–32× less data while often improving recall because you discarded noise.
-
Activation steering and interpretability. Collect residual-stream activations across many prompts, run SVD, and the top directions frequently correspond to human-interpretable concepts such as truthfulness, refusal, or code mode.
-
Low-rank adaptation (LoRA). When you train a LoRA adapter you are explicitly learning a low-rank update ( \Delta W = BA ). The mathematically optimal rank-( r ) factors for a given full update matrix are given by its truncated SVD. Training finds a nearby good factorization; the SVD view tells you the absolute best compression you could ever achieve for that rank.
-
Numerical hygiene in production. Any time you solve a linear system, compute an inverse, or even just do many dot products against a fixed matrix, a high condition number is a red flag. Monitoring the singular values (or at least the condition number of a small random projection) of your embedding or weight matrices is a cheap way to catch impending instability before it appears in your RAG quality metrics.
Practice problems
Do these with pencil and paper first, then verify with the code patterns above.
-
For the matrix ( A = \begin{pmatrix} 3 & 1 \ 1 & 3 \end{pmatrix} ), compute ( A^T A ), its eigenvalues, the singular values, and the condition number ( \kappa ). What changed compared with our first example?
-
Implement a tiny function
manual_svd_2x2(A)that accepts any 2×2 matrix, performs the eigendecomposition route, returns(U, S, Vt), and prints the reconstruction error. Test it on both matrices we used in the article. -
Generate (or hard-code) six 2-D points that form a clearly elongated cloud. Center them, run your manual SVD (or
np.linalg.svd), and print the first column of ( V ). Verify that it points roughly along the long axis of the cloud. -
Construct a deliberately ill-conditioned 5×3 matrix (make the third column almost equal to the sum of the first two plus a tiny ( \epsilon )). Add 0.001 noise to a single entry. Compute the change in the top singular vector before and after keeping only the largest two singular values. Quantify how much more stable the truncated version is.
Solution sketches
-
( A^T A = \begin{pmatrix} 10 & 6 \ 6 & 10 \end{pmatrix} ). Eigenvalues 16 and 4. Singular values 4 and 2. ( \kappa = 2 ). The matrix is better conditioned than the first example because the off-diagonal is relatively smaller.
-
The function is exactly the 10-line block shown in the “from-scratch” section. On both examples the max error is < 1e-14.
-
After centering, the first right singular vector will be within a few degrees of the direction of greatest elongation (you can eyeball it or compute the sample covariance and its dominant eigenvector - they must agree).
-
The full-rank top singular vector moves by a large angle when you perturb the near-dependent column. After truncation to rank 2 the direction is almost unchanged - the unstable direction has been removed.
Evaluation Rubric
- 1Carries out a complete SVD by hand for a 2×2 or 3×2 matrix with integer entries, computes singular values and vectors, and reconstructs the original matrix to verify A = UΣVᵀ
- 2Implements a from-scratch SVD routine using NumPy eigh on AᵀA, applies it to a toy embedding matrix, and contrasts reconstruction error and singular values against np.linalg.svd and sklearn TruncatedSVD
- 3Diagnoses a high-condition-number embedding matrix in a retrieval setting, explains how noise gets amplified in dot-product scores, and shows how keeping only the top-k singular components both compresses and denoises the representation
Common Pitfalls
- Forgetting to center (subtract column means) before treating a data matrix as input to PCA. The first principal component then points toward the data centroid instead of the direction of greatest spread.
- Running full np.linalg.svd on a 1-million-row embedding matrix when you only need the top 128 components; always use TruncatedSVD or randomized_svd for production scale.
- Reporting raw singular values as importance. Always convert to explained_variance_ratio = σ² / Σσ² so that you can say “the first 32 components explain 87 % of variance.”
- Treating U and V symmetrically. V lives in the original feature space (the columns you can interpret), U lives in the sample space (one coordinate per row of the original matrix).
- Applying SVD or PCA to unscaled features. A column with values in the thousands will dominate every singular vector even if it carries no semantic signal.
Follow-up Questions to Expect
Key Concepts Tested
singular value decomposition (SVD) hand computationeigendecomposition of AᵀA and relation to Vprincipal component analysis as SVD on centered datamatrix rank and truncated low-rank approximationcondition number and its effect on numerical stabilityapplications of SVD/PCA to embeddings, retrieval, and compression
Next Step
Next: Continue to Optimization Algorithms: Adam, Schedulers, and Why They Matter
You now own the factorizations that turn raw high-dimensional vectors into clean, stable, low-dimensional directions that embeddings and retrievers actually use. The optimization chapter shows you how those same matrix properties (condition numbers, curvature along the principal axes you just discovered) explain why plain gradient descent zig-zags or diverges, and why Adam plus learning-rate schedules succeed in practice. The two chapters together give you the complete mathematical story behind every training run.
References